Identifying Set-wise Differential Co-expression in Gene Expression Microarray Data
Sung Bum Cho, Jihun Kim and Ju Han Kim*
Background: Previous differential coexpression analyses focused on identification of differentially coexpressed gene pairs, revealing many insightful biological hypotheses. However, this method could not detect coexpression relationships between pairs of gene sets. Considering the success of many set-wise analysis methods for microarray data, a coexpression analysis based on gene sets may elucidate underlying biological processes provoked by the conditional changes. Here, we propose a differentially coexpressed gene sets (dCoxS) algorithm that identifies the differentially coexpressed gene set pairs between conditions.
Results: dCoxS is a two-step analysis method. In each condition, dCoxS measures the interaction score (IS), which represents the expression similarity between two gene sets using Renyi relative entropy. When estimating the relative entropy, multivariate kernel density estimation was used to model gene-gene correlation structure. Statistical tests for the conditional difference between the ISs determined the significance of differential coexpression of the gene set pair. Simulation studies supported that the IS is a representative measure of similarity between gene expression matrices. Single gene coexpression analysis of two publicly available microarray datasets detected no significant results. However, the dCoxS analysis of the datasets revealed differentially coexpressed gene set pairs related to the biological conditions of the datasets.

source code for computing interaction score (source.R)
simulation datasets I (.Rdata files)
SX; original data set
SX1; original data set + random values from N(0,0.01)
SX2; original data set + random values from N(0,0.5)
simulation datasets II (.Rdata files)
M; original data set
M1; original data set + random values
M2; original data set + random values
Raw expression plots and IS plots of SX,SX1 and SX2
source codes for computing and plotting differential co-expression of two gene sets (dcoxs.R, dcoxs.plot.R)
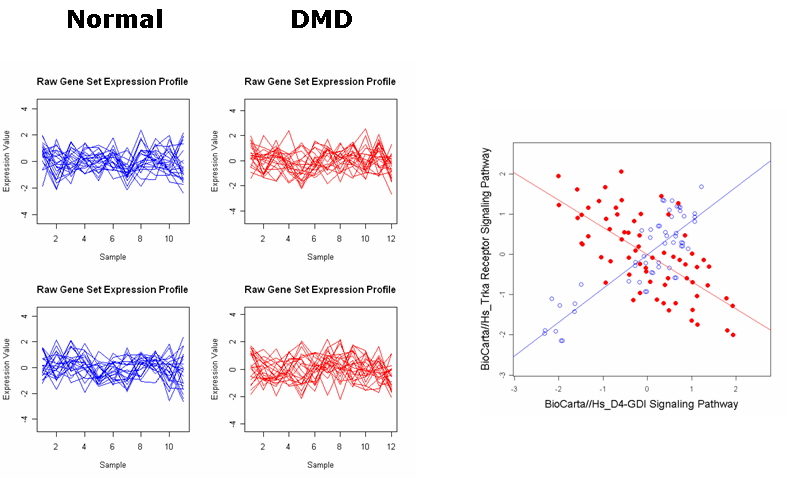
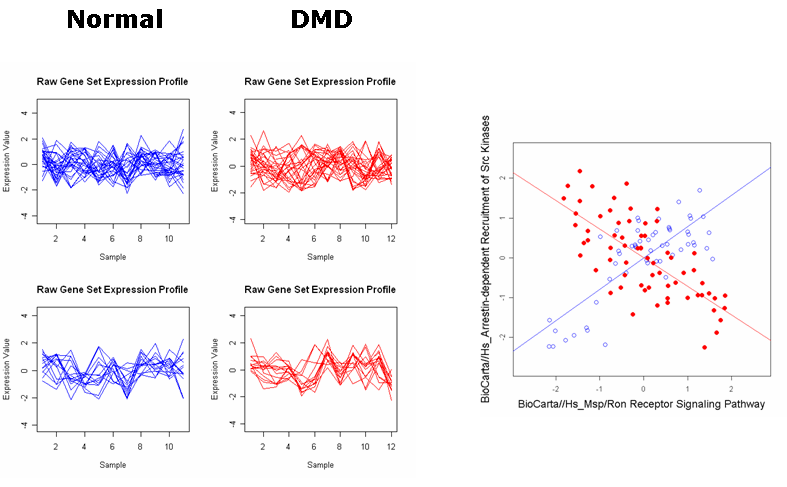
Raw expression profiles and IS plots in the Duchenne's muscular dystrophy (DMD) dataset.
1. Roles of arrestin-dependent Recruitment of Src Kinases in GPCR Signaling and Msp/Ron Receptor Signaling Pathway
2. Trka Receptor Signaling Pathway and D4-GDI Signaling Pathway