목차
Set-wise semantic similarity measures

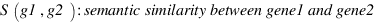

LordPW :
Semantic similarity measures as tools for exploring the gene ontology.
Lord PW, Stevens RD, Brass A, Goble CA
Pac Symp Biocomput() p601-12 (2003)
Lord PW, Stevens RD, Brass A, Goble CA
Pac Symp Biocomput() p601-12 (2003)
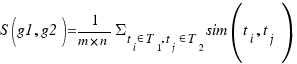
For this paper only those terms with evidence codes of “Traceable Author Statement” have been used.
FuSSiMeG :
Couto F, Silva M, Coutinho P
DI/FCUL TR 03--29 (2003 November)

Resnik, Jiang, and Lin measure :
Correlation between gene expression and GO semantic similarity.
Sevilla JL, Segura V, Podhorski A, Guruceaga E, Mato JM, Martínez-Cruz LA, Corrales FJ, Rubio A
IEEE/ACM Trans Comput Biol Bioinform2(4) p330-8 (2005 Oct-Dec) 10.1109/TCBB.2005.50
Sevilla JL, Segura V, Podhorski A, Guruceaga E, Mato JM, Martínez-Cruz LA, Corrales FJ, Rubio A
IEEE/ACM Trans Comput Biol Bioinform2(4) p330-8 (2005 Oct-Dec) 10.1109/TCBB.2005.50

RubioA :
Correlation between gene expression and GO semantic similarity.
Sevilla JL, Segura V, Podhorski A, Guruceaga E, Mato JM, Martínez-Cruz LA, Corrales FJ, Rubio A
IEEE/ACM Trans Comput Biol Bioinform2(4) p330-8 (2005 Oct-Dec) 10.1109/TCBB.2005.50
Sevilla JL, Segura V, Podhorski A, Guruceaga E, Mato JM, Martínez-Cruz LA, Corrales FJ, Rubio A
IEEE/ACM Trans Comput Biol Bioinform2(4) p330-8 (2005 Oct-Dec) 10.1109/TCBB.2005.50

LiebmanMN :
Assessing semantic similarity measures for the characterization of human regulatory pathways.
Guo X, Liu R, Shriver CD, Hu H, Liebman MN
Bioinformatics22(8) p967-73 (2006 Apr 15) 10.1093/bioinformatics/btl042
Guo X, Liu R, Shriver CD, Hu H, Liebman MN
Bioinformatics22(8) p967-73 (2006 Apr 15) 10.1093/bioinformatics/btl042

LinK
Prediction of yeast protein-protein interaction network: insights from the Gene Ontology and annotations.
Wu X, Zhu L, Guo J, Zhang DY, Lin K
Nucleic Acids Res34(7) p2137-50 (2006) 10.1093/nar/gkl219
Wu X, Zhu L, Guo J, Zhang DY, Lin K
Nucleic Acids Res34(7) p2137-50 (2006) 10.1093/nar/gkl219

funsim :
A new measure for functional similarity of gene products based on Gene Ontology.
Schlicker A, Domingues FS, Rahnenführer J, Lengauer T
BMC Bioinformatics7() p302 (2006 Jun 15) 10.1186/1471-2105-7-302
Schlicker A, Domingues FS, Rahnenführer J, Lengauer T
BMC Bioinformatics7() p302 (2006 Jun 15) 10.1186/1471-2105-7-302
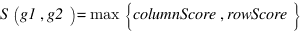
 ,
,
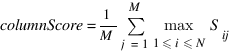
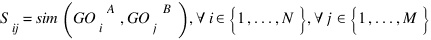
OlssonB :
Combining functional and topological properties to identify core modules in protein interaction networks.
Lubovac Z, Gamalielsson J, Olsson B
Proteins64(4) p948-59 (2006 Sep 1) 10.1002/prot.21071
Lubovac Z, Gamalielsson J, Olsson B
Proteins64(4) p948-59 (2006 Sep 1) 10.1002/prot.21071

GORank :
김기성, 유상원, 김형주
정보과학회논문지 33권 7호 p682-692 (2006 December)


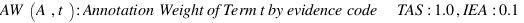

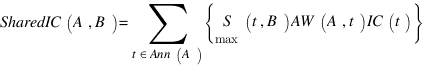
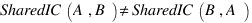

LussierYA :
Evaluation of high-throughput functional categorization of human disease genes.
Chen JL, Liu Y, Sam LT, Li J, Lussier YA
BMC Bioinformatics8 Suppl 3() pS7 (2007 May 9) 10.1186/1471-2105-8-S3-S7
Chen JL, Liu Y, Sam LT, Li J, Lussier YA
BMC Bioinformatics8 Suppl 3() pS7 (2007 May 9) 10.1186/1471-2105-8-S3-S7

G-SESAME :
A new method to measure the semantic similarity of GO terms.
Wang JZ, Du Z, Payattakool R, Yu PS, Chen CF
Bioinformatics23(10) p1274-81 (2007 May 15) 10.1093/bioinformatics/btm087
Wang JZ, Du Z, Payattakool R, Yu PS, Chen CF
Bioinformatics23(10) p1274-81 (2007 May 15) 10.1093/bioinformatics/btm087





LussierYA :
Information theory applied to the sparse gene ontology annotation network to predict novel gene function.
Tao Y, Sam L, Li J, Friedman C, Lussier YA
Bioinformatics23(13) pi529-38 (2007 Jul 1) 10.1093/bioinformatics/btm195
Tao Y, Sam L, Li J, Friedman C, Lussier YA
Bioinformatics23(13) pi529-38 (2007 Jul 1) 10.1093/bioinformatics/btm195

BurgunA :
A transversal approach to predict gene product networks from ontology-based similarity.
Chabalier J, Mosser J, Burgun A
BMC Bioinformatics8() p235 (2007 Jul 2) 10.1186/1471-2105-8-235
Chabalier J, Mosser J, Burgun A
BMC Bioinformatics8() p235 (2007 Jul 2) 10.1186/1471-2105-8-235
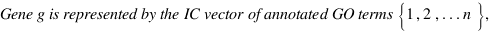
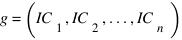

DongXu :
Quantitative assessment of relationship between sequence similarity and function similarity.
Joshi T, Xu D
BMC Genomics8() p222 (2007 Jul 9) 10.1186/1471-2164-8-222
Joshi T, Xu D
BMC Genomics8() p222 (2007 Jul 9) 10.1186/1471-2164-8-222
Example of GO index and the corresponding GO ID and functional category
| Index level | GO Index | Functional category and GO ID |
|---|---|---|
| Index1 | 1-2 | cellular process (GO:0009987) |
| Index2 | 1-2-1 | cell communication (GO:0007154) |
| Index3 | 1-2-1-8 | signal transduction (GO:0007165) |
| Index4 | 1-2-1-8-1 | cell surface receptor linked signal transduction (GO:0007166) |
| Index5 | 1-2-1-8-1-4 | G-protein coupled receptor protein signaling pathway (GO:0030454) |
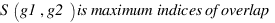


ZhangA :
Semantic integration to identify overlapping functional modules in protein interaction networks.
Cho YR, Hwang W, Ramanathan M, Zhang A
BMC Bioinformatics8() p265 (2007 Jul 24) 10.1186/1471-2105-8-265
Cho YR, Hwang W, Ramanathan M, Zhang A
BMC Bioinformatics8() p265 (2007 Jul 24) 10.1186/1471-2105-8-265
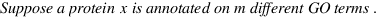
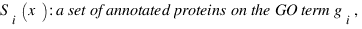
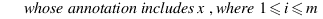
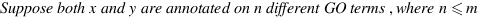
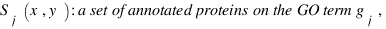
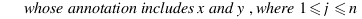
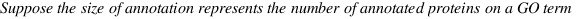
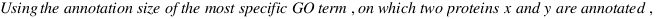
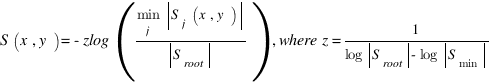
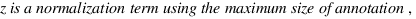
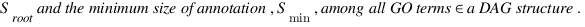
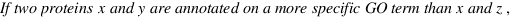
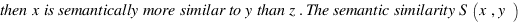

simGIC :
Metrics for GO based protein semantic similarity: a systematic evaluation.
Pesquita C, Faria D, Bastos H, Ferreira AE, Falcão AO, Couto FM
BMC Bioinformatics9 Suppl 5() pS4 (2008 Apr 29) 10.1186/1471-2105-9-S5-S4
Pesquita C, Faria D, Bastos H, Ferreira AE, Falcão AO, Couto FM
BMC Bioinformatics9 Suppl 5() pS4 (2008 Apr 29) 10.1186/1471-2105-9-S5-S4
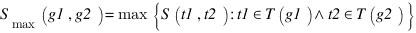
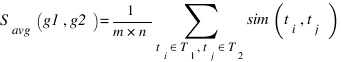
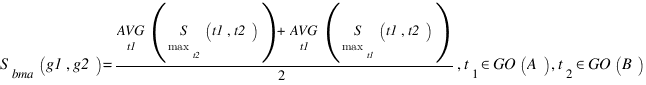
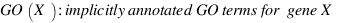
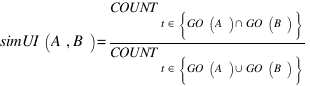

GOSAP :
GOSAP: Gene Ontology-Based Semantic Alignment of Biological Pathways.
Gamalielsson J, Olsson B
Int J Bioinform Res Appl4(3) p274-94 (2008)
Gamalielsson J, Olsson B
Int J Bioinform Res Appl4(3) p274-94 (2008)