Semantic similarity measures
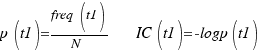

Resnik, Lin, Jiang&Conrath :
P. Resnik Using Information Content to Evaluate Semantic Similarity in a Taxonomy Proc. 14th Int’l Joint Conf. Artificial Intelligence, pp. 448-453, 1995.
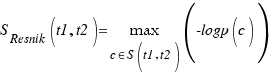
A drawback of the Resnik measure is that it does not differentiate between two terms if their subsumer is the same.
D. Lin An Information-Theoretic Definition of Similarity Proc. 15th Int’l Conf. Machine Learning, pp. 296-304, 1998.
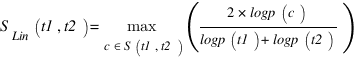
J.J. Jiang and D.W. Conrath Semantic Similarity Based on Corpus Statistics and Lexical Taxonomy Proc. Int’l Conf. Research in Computational Linguistics, ROCLING X, 1997.

As we argued with respect to Jiang, we encounter an important drawback when both gene products happen to be shallowly annotated: similarity values would appear to be deceptively high, while this might not in fact be true. For example, if we had two genes annotated by the term “intracellular,” (GO:0005622), and no additional annotation was available, Jiang distance will be nil, and Lin similarity will be one. Although both measures provide excellent similarity results, in reality, both gene products are likely to be quite different.
Schlicker :
Schlicker A, Domingues FS, Rahnenführer J, Lengauer T
BMC Bioinformatics7() p302 (2006 Jun 15) 10.1186/1471-2105-7-302

Schlicker A, Rahnenführer J, Albrecht M, Lengauer T, Domingues FS
Genome Biol8(3) pR33 (2007) 10.1186/gb-2007-8-3-r33
Schlicker A, Albrecht M
Nucleic Acids Res36(Database issue) pD434-9 (2008 Jan) 10.1093/nar/gkm806
Schlicker A, Albrecht M
Nucleic Acids Res38(Database issue) pD244-8 (2010 Jan) 10.1093/nar/gkp979
Schlicker A, Lengauer T, Albrecht M
Bioinformatics26(18) pi561-7 (2010 Sep 15) 10.1093/bioinformatics/btq384
LinK :
Wu X, Zhu L, Guo J, Zhang DY, Lin K
Nucleic Acids Res34(7) p2137-50 (2006) 10.1093/nar/gkl219
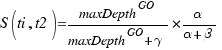
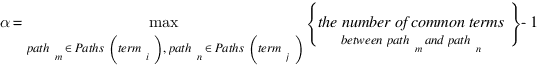
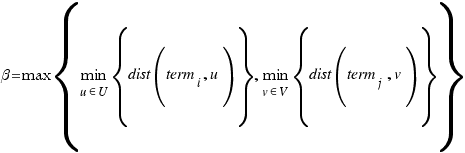
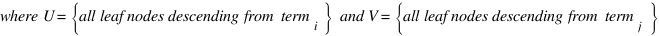

CoutinhoP
Couto F, Silva M, Coutinho P
Data & Knowledge Engineering 61 p137-152 (2007 April)
Example GO tree
 is a - 0.8, part of - 0.6
is a - 0.8, part of - 0.6
S-values for GO terms in DAG for term Intracellular membrane-bound organelle:0043231
| GO terms | S-value |
|---|---|
| 43231 | 1.0 |
| 43229 | 0.8 |
| 43227 | 0.8 |
| 5622 | 0.48 |
| 5623 | 0.288 |
| 43226 | 0.64 |
| 5575 | 0.512 |
S-values for GO terms in DAG for term Intracellular organelle:0043229
| GO terms | S-value |
|---|---|
| 43229 | 1.0 |
| 5622 | 0.6 |
| 5623 | 0.36 |
| 43226 | 0.8 |
| 5575 | 0.64 |
Semantic similarity between GO term A and B:

LussierYA :
Tao Y, Sam L, Li J, Friedman C, Lussier YA
Bioinformatics23(13) pi529-38 (2007 Jul 1) 10.1093/bioinformatics/btm195
use Lin's semantic similarity, but use occurrence probability of a term

Yang & Kim :

Meeta Mistry :
Mistry M, Pavlidis P
BMC Bioinformatics9() p327 (2008 Aug 4) 10.1186/1471-2105-9-327
TO : Term Overlap
 : set of all direct annotations for each gene and all of their associated parent terms (excluding root)
: set of all direct annotations for each gene and all of their associated parent terms (excluding root)

Sidahmed Benabderrahmane :
Benabderrahmane S, Smail-Tabbone M, Poch O, Napoli A, Devignes MD
BMC Bioinformatics11(1) p588 (2010 Dec 1) 10.1186/1471-2105-11-588
Jain S :
Jain S, Bader GD
BMC Bioinformatics11(1) p562 (2010 Nov 15) 10.1186/1471-2105-11-562
Wang H :
Wang H, Zheng H, Azuaje F
Bioinformatics26(20) p2643-4 (2010 Oct 15) 10.1093/bioinformatics/btq477
Zhu W :
Zhu W, Hou J, Chen YP
J Theor Biol267(2) p129-36 (2010 Nov 21) 10.1016/j.jtbi.2010.08.005
Tedder PM :
Tedder PM, Bradford JR, Needham CJ, McConkey GA, Bulpitt AJ, Westhead DR
Bioinformatics26(19) p2431-7 (2010 Oct 1) 10.1093/bioinformatics/btq450
Chen Z :
Chen Z, Tang J
Int J Data Min Bioinform4(5) p520-34 (2010)
Wang Z :
Wang J, Zhou X, Zhu J, Zhou C, Guo Z
BMC Bioinformatics11() p290 (2010 May 28) 10.1186/1471-2105-11-290
Sheehan B :
Sheehan B, Quigley A, Gaudin B, Dobson S
BMC Bioinformatics9() p468 (2008 Nov 4) 10.1186/1471-2105-9-468